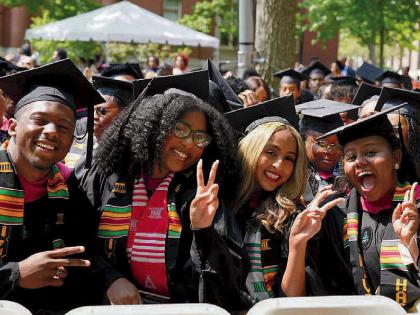When George McDonald Church arrived at Harvard in 1977, nine out of 10 biologists did research without touching a computer. They wrote journal articles on IBM typewriters, not word processors, and used slide rules and portable calculators to analyze data. Even recombinant DNA technology, which empowered biologists to cut and splice DNA and engineer genes, was artisanal. The small coterie of computer-savvy biologists were x-ray crystallographers, who fired beams of radiation at specially prepared samples to discover the shapes of life's essential molecules. These experiments generated so much raw data that computers were needed to extract clear structural images.
 |
| Away from his laboratory, George Church often works beside his backyard fish pond. With daughter Marie, he explores a different kind of life science: breeding turtles. |
| Photograph by John Soares |
Into this world burst Church, an energetic 23-year-old who had graduated from Duke University three years earlier, entered graduate biochemistry and virology programs there immediately, and flunked out of both. Since childhood, he had been caught up in computers, chemistry, mathematics, and biology; in graduate school he realized that all four disciplines converged in x-ray crystallography. So smitten was Church, so passionately engaged in this new world, that he pretty much forgot about his classes. Although Duke dismissed him, he published five scientific articles about what he had learned in the lab, an extraordinary and precocious accomplishment. Duke's loss was Harvard's gain: Church was admitted to the Faculty of Arts and Sciences (FAS) doctoral program in biochemistry and molecular biology.
Twenty-six years later, Church, Ph.D. '84, is a professor of genetics at Harvard Medical School (HMS) and director of the Lipper Center for Computational Genetics. He runs one of the largest research labs in the Longwood Medical area, a thrumming engine of productivity with 40 people and more than twice as many computers. About half of those people work primarily in the "wet lab," performing experiments to answer such basic questions as why cancer cells proliferate faster than normal ones. Using DNA microarrays, a popular and versatile tool that can monitor the activities of thousands of genes at once, they have been able to create "fingerprints" for malignant cells showing which genes are turned on as the cells divide. Experimental treat normal ones. Using DNA microarrays, a popular and versatile tool that can monitor the activities of thousands of genes at once, they have been able to create "fingerprints" for malignant cells showing which genes are turned on as the cells divide. Experimental treatments that act on proteins made by these genes are being tested on patients by doctors at other institutions.
Computation and experimental biology are inseparable in the Church lab. Graduate students routinely complete an experiment at the bench, strip off their latex gloves, and sit down at the computer to tweak elaborate mathematical models incorporating their latest results. The model then helps them decide what the next experiment should be.
The other wing of Church's large, L-shaped suite in the medical school's New Research Building (NRB) is occupied by Lipper Center scientists, who are teasingly called "propeller heads" by their wet-lab colleagues. The Lipper team helps other Harvard researchers design high-throughput experiments, such as those using microarrays or automated analyzers that evaluate hundreds of potential drugs in a single run. Tools like these can generate 100,000 facts before lunch. When Church was in graduate school, it sometimes took years to identify genetic abnormalities linked to specific human diseases such as cystic fibrosis or muscular dystrophy. Such discoveries inevitably raised hopes that an effective treatment was just around the corner, but all too often potential drugs interfered with the gene's product yet did nothing to combat the disease, forcing researchers to resume the search for additional targets for treatment. Now that microarrays and computers can capture the activity of thousands of genes simultaneously, Church says, serendipitous discoveries of multiple drug targetsand drugs that hit the bull's-eyeare far likelier to emerge.
The Lipper Center serves a second purpose as well. When medical-area researchers need a new technology that doesn't yet exist, or are uneasy because a new machine doesn't seem to work quite right, they often turn to the Lipper Center for help. For example, many researchers clamored for a microarray with the whole genome of Escherichia coli, the bacterium that is widely used in biotechnology and all sorts of lab research. Church's group encouraged California-based Affymetrix Inc. to create such a chip, helped design it, and developed methods for using it and software to help researchers interpret their results. In collaboration with Aventis, a major pharmaceutical company, this chip has been used in the Church lab to understand in detail how a new class of antibiotics works to kill bacteria. Without the chip, the researchers would not have been able to see the drug's many different effects. The propeller heads have also been test pilots for countless new computers and for huge robotic analyzers that cost millions of dollars.
Until the New Research Building opened this past fall, people from the wet lab and the Lipper Center met together weekly but spent the rest of the time on opposite sides of the HMS Quad. Now that they're finally under one roof, Church expects more people to be caught up in the kind of design and consulting work that the center has been doing. When he brought everyone together for the first meeting at the NRB, the 30 lab members in attendance seemed still to be settling in, much like college students who've moved in with new roommates. Someone said that the "computational people" were worried because they didn't know what to do when the alarm on a piece of lab equipment rangwhich happens fairly often.
"Run," Church joked. "Run as fast as you can." There was a big laugh and the group loosened up. He introduced the day's speaker, a biophysics graduate student in the lab who described modeling yeast metabolism using about 1,000 variables. Church sat alone in the front row, long legs stretched out in front of him, occasionally tipping his chair onto its back legs when something caught his attention. (Church has what he describes as "a touch of narcolepsy," and one former graduate student recalls being mid-way though a similar presentation and thinking with horror, "Oh my God, I've put him to sleep." Suddenly Church's head snapped up and he said, "You missed a minus sign in the third term of your equation.")
Church's ability to bring together information technology and experimental genetics has made him a "force majeure in science," according to Philip Leder, Andrus professor of genetics and head of the genetics department at HMS. Far from being "just a computer geek," Leder says, Church is a polymath who "has terrific ideas that nobody else would think of putting together, because of the many disciplines he has mastered." A graduate of the lab goes even further. "He is regarded with awe by everybody I've ever met," says Douglas Selinger, Ph.D. '02, a bioinformatics scientist at Novartis who did his doctoral research under Church's guidance.
"He's always working right on the edge of what's possible," observes Martha Bulyk, who completed her doctorate in biophysics in the Church lab in 2000. After an abbreviated, nine-month postdoctoral fellowship there, she was hired as an assistant professor of genetics at HMS and Brigham and Women's Hospitalone of 18 former lab members who've been snapped up by leading academic institutions.
Church is a tall, physically imposing man whose intense blue-green eyes, full beard, and craggy brow give him a touch of the Old Testament prophet. Yet there is nothing intimidating about his low-key, amiable manner. Young scientists who have trained in his lab describe their mentor as "a great guy" as well as a visionary thinker.
One person's visionary is another's oddball, of course. In 1994, then first-year graduate student Preston "Pete" Estep, Ph.D. '00, asked an administrator where he could learn about cutting-edge applications of computer science to biology. "There's this guy named George Church, and if he's not weird enough for you, then there's nothing for you at Harvard," the answer came back. "This is the most out-there stuff that we've got."
Church's latest vision is that society is about to be transformed by a genetics revolution that will echo what has already happened with computers. Twenty years ago, relatively few people had a PC or an Apple. Now it's hard to find a household without a computer or at least an appliance with a rudimentary brain, or to find a person who hasn't looked up a flight schedule or bought a sweater on the Web. "Computer power is dirt cheap and it's almost user-friendly," Church says with a laugh.
He believes genetics is on a similar trajectory. Steady improvements in automated sequencing technologies enabled scientists to complete the human genome project in 2001, and to sequence the entire genetic complements of certain yeasts, bacteria, viruses, worms, mice, and a growing list of larger creatures, including dogs and chimpanzees. These accomplishments, he says, now put genetics "where computing was just before the World Wide Web."
The next step, as Church has been telling interviewers from CNN, MSNBC, U.S. News & World Report, and the Wall Street Journal, among others, is for individuals to have their personal genomes sequenced. Church is not a publicity seeker by nature, but he has been on the stump about personal genomes because he believes that if the idea catches fire, the benefits for individual health and biomedical research will be enormous. For example, young people who dismiss generic advice about healthy eating will be far more likely to listen if their doctor says they have a specific gene that dramatically increases their risk of heart disease. And if enough people make their genomes available for study, researchers should be able to design better drugs for hard-to-treat diseases like Alzheimer's and figure out why a medication helps one person but not another.
"I am convinced that we will want our personal genome possibly more than we want a personal computer," Church says. Although obtaining such information is extremely expensive at present, he anticipates that "the same kind of people who pay to go into space" will soon be having their genetic code deciphered. As for making personal genomes as affordable as personal computers, he says, "I don't know what year that will be, but it will probably happen so fast that people will be amazed."
Not surprisingly, he has his own ideas about how to increase the quality and speed of DNA sequencing while cutting costs. "For 20 years, the central problem that George has grappled with is the speed of DNA sequencing," says Pete Estep, who is now chief executive officer of Longenity, a Boston-based biotechnology company that is working on anti-aging strategies. (Estep and other former Church lab members have started eight biotech companies since the late 1980s, and six of the researchers are presently CEOs. Estep attributes this in large part to the example Church sets for his students: "It's understood in the lab that there's this fantastically exciting opportunity, and if you don't grab on, you're gonna get left behind." Church has given a helping hand to entrepreneurial former students and serves on the scientific advisory boards for most of the companies they've founded.)
Church says he works mainly to lower the cost and improve the quality of sequencing, and feels that "speed takes care of itself" if you focus on these. He believes that better and faster sequencing will bring many benefits: the ability to sequence the genomes of many species offers academic biologists a window on evolution, for example, and access to a patient's genome will help doctors predict risk, diagnose disease, and choose the treatment most likely to help without causing adverse effects. Church has therefore invented a series of methods for automating the reading of genetic code, each better than its predecessor. He calls the most recent method "polony" technology (see "Polony Power"); although many academic and private teams are working to develop fast, affordable sequencing technologies, this is one of the swiftest in the race. Before it can be transferred from the research lab to the clinic, however, or be used to sequence personal genomes en masse, Church says his group needs to bring down the cost and make the procedure foolproof.
There are ethical and social concerns as well. Once a new personal genome is assembled, Church notes, computer comparisons can detect differences between it and other genomes, "some of which are going to be innocuous and others alarming." This could exacerbate an issue that arose when it first became possible to screen for diseases such as Huntington's and breast cancer: many people from affected families refuse testing because they worry about losing their jobs or their health insurance. By the time personal genomes become affordable, however, new legal protections may have resolved this problem: a bill prohibiting use of genetic data in hiring or insurance decisions passed the U.S. Senate this past October, but at press time had not yet been scheduled for hearings in the House. In the long run, Church says, genetic discrimination will be pointless because every individual's genome will reveal vulnerability to some health problem.
As compelling as Church finds the idea of the personal genome, it is only one element in a sprawling research portfolio. On the medical front, his lab collaborates with scientists at several Harvard-affiliated hospitals to study cardiovascular disease, traumatic injury, and stress responses. For example, in a collaboration with Penney professor of neurology Bradley T. Hyman at Massachusetts General Hospital, the Church group is studying genetic factors that predispose people to develop Alzheimer's disease, looking for mechanisms of action that might someday be blocked by therapeutic drugs.
At the other end of the spectrum, he is involved in research on environmental remediation: along with colleagues at MIT and two Harvard hospitals, he has received $15 million from the Department of Energy to study some microbes that remove toxic substances from the ocean and others that lower carbon dioxide levels in the atmosphere. He's also waiting to hear about a grant that would give him the resources to pursue one of his newer passions, the idea of reprogramming adult stem cells to cure all sorts of illnesses.
Highly motivated graduate students from molecular biology, chemistry, microbiology, physics, computer science, and other disciplines are drawn into Church's orbit. When Rob Mitra interviewed for a place in the lab in 1997, Church told him, "I believe that what we're doing is the most important thing that anyone could possibly be doing, and I'm looking for people who believe that as well." Mitra was hooked. Although he arrived with an electrical-engineering background, he learned enough biology to share one of the polony-related patents with Church. Says Mitra, who is now an assistant professor of genetics at Washington University in St. Louis, "It's sort of a Renaissance lab, because he is sort of a Renaissance professor."
Like any true Renaissance figure, Church has a domestic as well as a public side. Everyone who comes to know him soon realizes how important his family is to him. He fell for geneticist Chao-ting Wu '76, Ph.D. '85, when they were in graduate school together at Harvard, and followed her to California when she landed a postdoc at Stanford University. They came back east two years later, mostly on her initiative, and were married in 1990. Today, Ting Wu is an associate professor of genetics at HMS and the couple have a 12-year-old daughter, Marie. The family lives within walking distance of the medical area, and when Marie comes home from school, Church is often beside the fish pond in the garden, writing an article or a grant proposal. For the past two years, father and daughter have shared the accomplishment of breeding rare hingeback turtles, a process that involves incubating eggs under carefully controlled conditions.
Looking back, Church says he was hesitant to start either a lab or a family, because he had seen dysfunctional versions of each. "But I lucked out with both my child and my lab, and they are wonderful, functioning, happy people and places."
Everything that happens in the Church lab today reflects the man and his history. Growing up in Clearwater, Florida, the only way the 10-year-old Church could get his hands on a computer was to build one with components from an electrical-supply store. Soon after that, his adoptive father, a physician educated in New England, sent young George to his alma mater, Phillips Academy in Andover. Church still beams when he recalls arriving there: "It was like walking into Oz, out of black and white and into color." Finally he had access to a real computer, via a remote hookup to a GE-635 mainframe at Dartmouth College. (Although he'll always have a soft spot for this, his first computer, he notes that those he uses today run about one million times faster.)
As an undergraduate at Duke, Church had an epiphany: DNA, which stores instructions for making all living creatures, is a stream of information much like the linear programs he was learning to write. The idea of using manufactured computers to analyze biological ones was born then, and is still growing as he approaches his fiftieth birthday.
"Certain people have things in them that have to come out," Church muses during a conversation in his glass-walled office. "I've felt since I was young that the computer thingand now this personal genome ideaare visions that are burning inside. You put together little crude things, trying to get what's outside to match what's inside. And you keep going because the images are in there and you are driven to do it. The first computer that I made didn't satisfy me, but it satisfied some urge. I'm still looking for a computer that is intelligent, that knows more than I know. And it's getting closer."
Although the young scientists who have been mentored by Church may not have heard him utter these exact words, they know what's going on. "Some people do things for money, and some people do things for the joy of knowing," says Pete Estep. "George is 100 percent in the 'joy of knowing' camp."
Patricia Thomas has written recently for Harvard Magazine about neuroscience and research on infection.